Guest researcher blog post about the recently published paper on the identification of potential new biomarkers in the blood, in the journal Brain Communications. Written by lead author, Dr Greig Joilin, Postdoctoral Research Fellow at the University of Sussex:
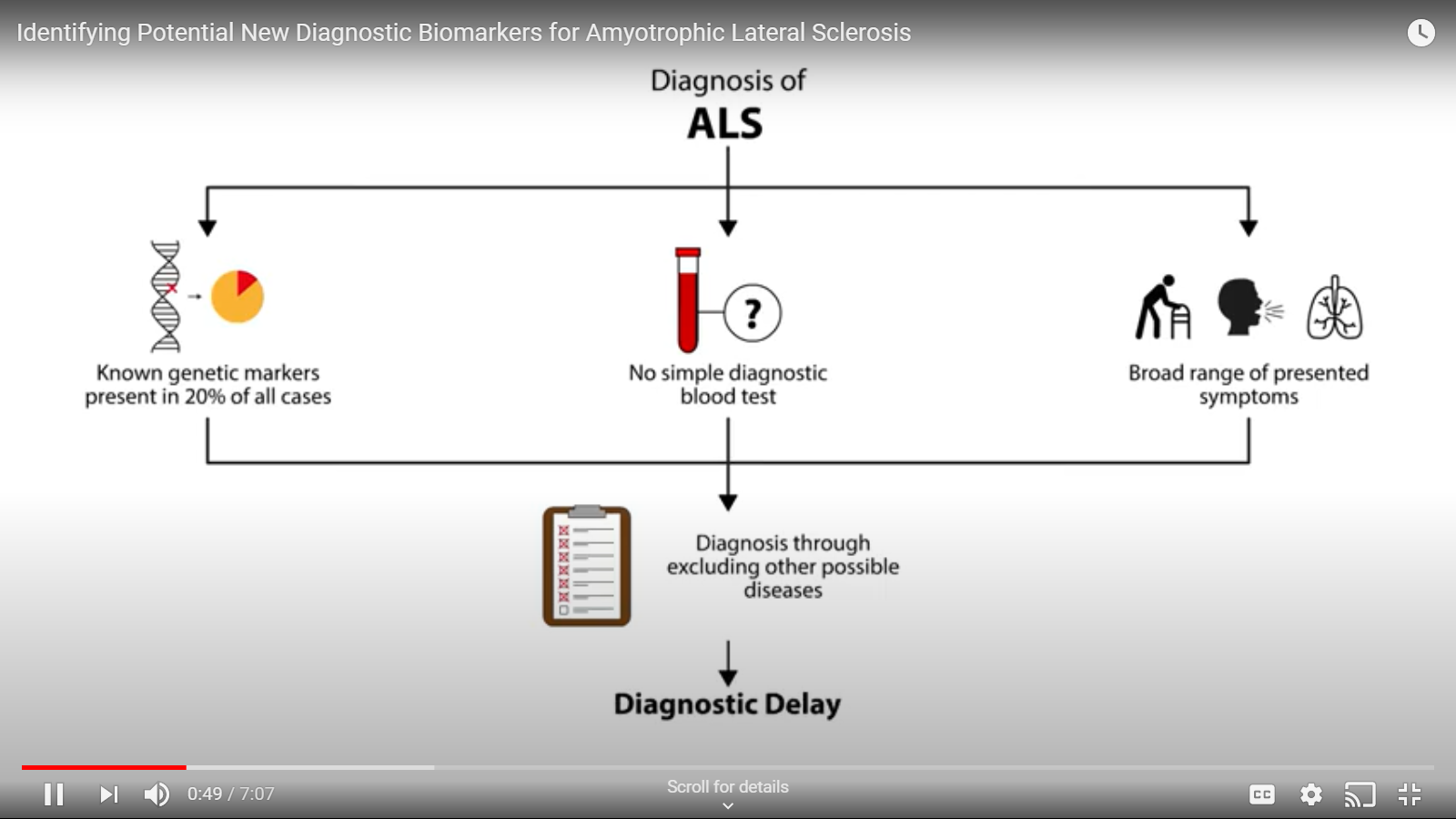
Diagnosis of MND is currently a time-consuming process of elimination, causing patients and families prolonged stress and uncertainty, as well as delaying potential treatment. Finding clear molecular biomarkers that can aid in the diagnostic process and predicting prognosis (the likely course of disease) is important for improving quality of life and the development of drugs. Through a dedicated collaboration of clinicians and scientists, and by using samples generously provided by people affected by MND, we have taken some important first steps towards improving this situation. Here we have identified seven non-coding RNA molecules that can be used to distinguish MND blood samples from healthy controls. We’ve created an animation (above) that takes you through our project and our results, and expanded on this below.
Firstly, who am I?
 My name is Greig Joilin and it is a privilege to share the work that I have primarily been undertaking at the University of Sussex in Brighton. I completed my PhD at the University of Otago in Dunedin, New Zealand, looking at what changes occur to help form memories. When I finished, I wanted to be able to apply my skills to research that had more clinical outcomes. Therefore, I came over to the UK to work on this project as I was excited about the possibility of helping to identify RNA-based biomarkers that could help improve the diagnostic delay, and the potential implications this might have on understanding the pathology of the disease on a molecular level. Since 2016, I have been working on this project and we now have some exciting results to share.
My name is Greig Joilin and it is a privilege to share the work that I have primarily been undertaking at the University of Sussex in Brighton. I completed my PhD at the University of Otago in Dunedin, New Zealand, looking at what changes occur to help form memories. When I finished, I wanted to be able to apply my skills to research that had more clinical outcomes. Therefore, I came over to the UK to work on this project as I was excited about the possibility of helping to identify RNA-based biomarkers that could help improve the diagnostic delay, and the potential implications this might have on understanding the pathology of the disease on a molecular level. Since 2016, I have been working on this project and we now have some exciting results to share.
What are we looking for?
Over the last four years with the support of the Motor Neurone Disease Association, as part of a collaboration led by Professor Majid Hafezparast with Professors Nigel Leigh, Sarah Newbury and Martin Turner, we have been working on discovering biomarkers to help with the diagnosis of MND. Biomarkers are any measurable indicator of a biological change, but it is often used in relation to disease, and so we are looking for something that changes in relation to MND. While biomarkers can include changes in brain structure as measured by imaging studies for example, we are interested in identifying a molecular biomarker as these are relatively less invasive to measure.
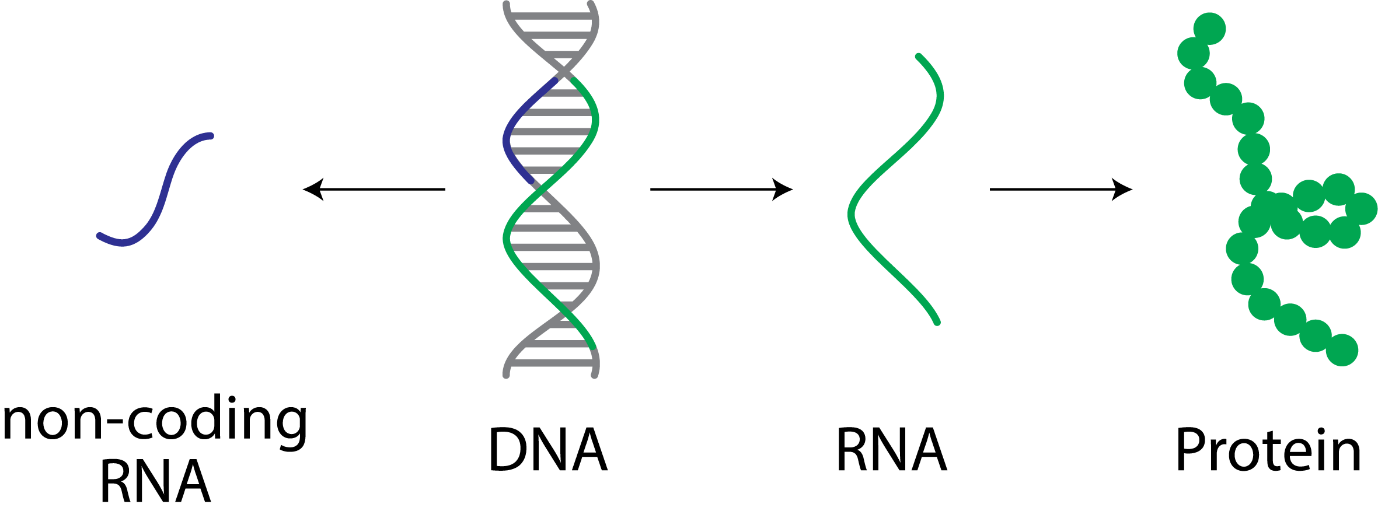
Our particular molecules of interest are called RNA. While DNA is well-known as containing our genes, or the instructions that makes us who we are, RNA is the molecule that allows these instructions to be read, and for them to be expressed as proteins. Without RNA, the information in our genes simply do not get expressed. While these RNA help code for proteins, there are some RNA which do not code for proteins, called non-coding RNA (ncRNA). Some of these are important for regulating the amount of proteins in the cell. Interestingly though, these ncRNA can also travel between cells through the blood, allowing for cells to communicate between each other across the body and regulate protein expression in other cells, and we believed that these ncRNA molecules would change in diseases such as MND.
So, we have been examining serum, a compartment of blood without all the red and white blood cells, to look for ncRNA-based biomarkers. For our study, we were able to use 79 samples of blood that were generously donated by volunteers and collected by our collaborator Professor Martin Turner as part of The Oxford Study for Biomarkers in MND (BioMOx) which includes people with MND or diseases similar to MND, and healthy people.
Previous studies investigating ncRNA-based biomarkers focused on the small subclass called microRNA, but our novel approach was to broaden this to include all small ncRNA to increase the range of potential candidates. Using next generation sequencing, technology which has revolutionised genomic research and allowed us to easily profile all ncRNA across multiple samples at the same time, we identified several potential candidates and confirmed dysregulation of seven ncRNA between MND patients and healthy controls.
Are these biomarkers reliable for use?
One ncRNA of particular note was hsa-miR-206, which has been previously identified as a potential biomarker for MND. We were able to show that it was mainly detected in MND patients and not in healthy people. However, it was also detected in patients with diseases similar to MND, so like neurofilaments, hsa-miR-206 is not specific to the disease.
Indeed, one of the main problems with any biomarker discovery project is whether the candidate biomarker is specific for MND, and so relying on one biomarker increases its chances of not being specific. Of course, using multiple biomarkers is a good idea, but how to use them together is something that needs to be considered.
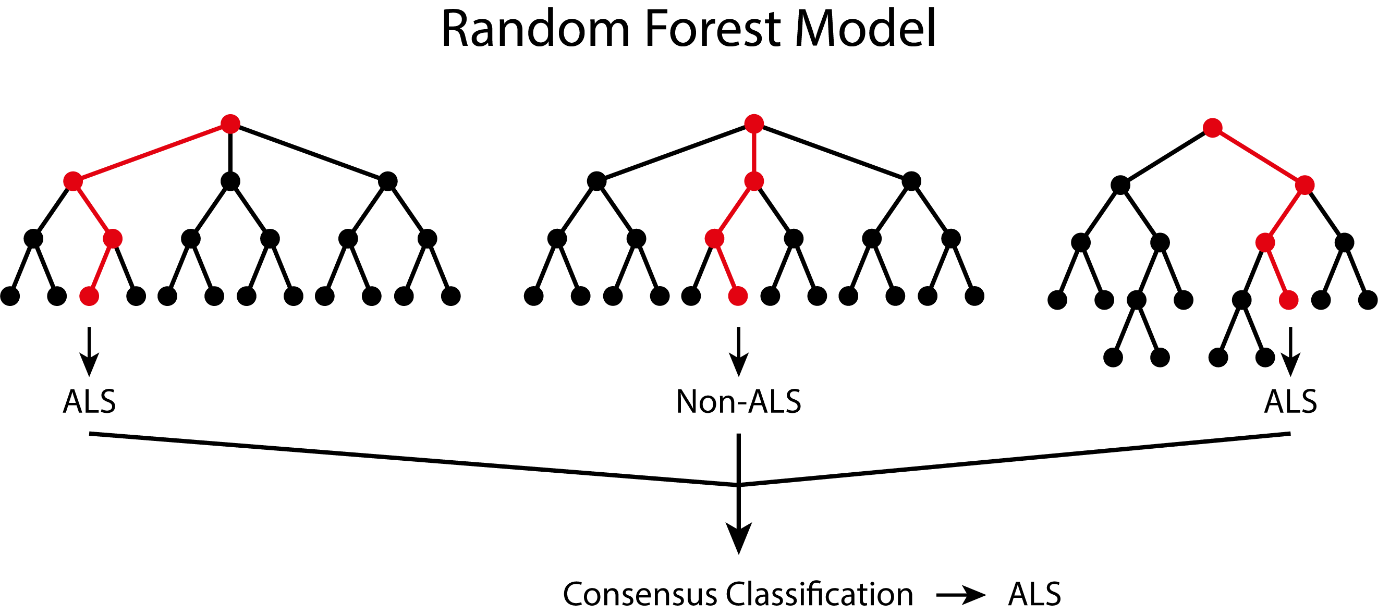
For our study, we took our biomarker candidates as a group and used their expression in a unique way by utilising a statistical model called a random forest model. Through using multiple decision trees, this model classifies the samples as either being from a person with MND or not, as illustrated in our animation.
Using the BioMOx samples, we were able to get an accuracy of nearly 94% for assigning a sample being MND or non-MND, which is very encouraging. To investigate whether these biomarker candidates and the model were specific to these samples, we obtained further samples from The ALS Biomarker Study run by Professor Andrea Malaspina at the Queen Mary University of London and the Ulm Neurological Biobank run by Professor Albert Ludolph in Germany. Using these additional 157 samples, we were able to classify these samples with a 74% accuracy using the same biomarkers and model.
What comes next?
We believe that this is highly encouraging and is an important first step to aid with identifying an RNA-based biomarker signature for diagnosing MND. It is important to note that at this point, this biomarker signature is not yet ready to be used clinically. We believe that we need to work to improve its accuracy further across groups of MND patients and healthy controls from different countries to help introduce greater flexibility in the signature. We wish to develop the signature further so that it may be able to provide an early indication about disease progression. Nonetheless, these goals are feasible, and it is definitely possible that this may be used to help diagnose MND patients in the future.
Moving forward, I also hope to begin work to identify further ncRNA biomarkers to help aid in predicting prognosis and look at how these biomarkers change across disease progression. Additionally, I want to investigate the expression of ncRNA in the cerebrospinal fluid (liquid surrounding in the brain and spinal cord) of MND patients and compare with findings observed in serum, as this may give us an indication as to what the causes of these alterations in the blood. While this work was suspended with the lockdown due to the COVID-19 pandemic, restrictions have begun to ease at the University of Sussex, and we have begun to restart this work. While initially we will be limited in what we can do, we are excited to be back in the lab and test new potential targets that we identified during the lockdown in both serum and cerebrospinal fluid.
I want to take this opportunity to thank all our collaborators across the UK and Germany for all their help with this work. I also want to thank the MND Association who have funded this project for the last four years and without whom we would not have been able to generate these crucial first steps. Lastly, and most importantly, I want to thank all the people who donated samples for this work, as none of this would be possible without their collaboration and generosity.
If you’d like to keep updated on our research or would like to contact us, please follow us on Twitter: @DrGreigJoilin and the lab @HafezparastLab.
Many thanks to Greig and the team for writing this blog post and for creating such an informative animation to help illustrate this important research.
- Read the full article in Brain Communications
- Watch Greig’s animation on YouTube
- Read about the research we fund in 2020 here.